|
Size: 1165
Comment:
|
Size: 2495
Comment:
|
| Deletions are marked like this. | Additions are marked like this. |
| Line 1: | Line 1: |
| == EMAN2 alpha == Note that EMAN2 is still in alpha-testing. It can do some useful things, but is not completely stable yet. |
== EMAN2 == EMAN2 is now stable enough to be a complete replacement for EMAN1 for most tasks. It is capable of single particle reconstructions at near-atomic resolution, and can process very large data sets (>100,000 particles). |
| Line 4: | Line 5: |
| * [:EMAN2/Install:Installation Instructions and Tips] * [:EMAN2/Programs:Individual Program Documentation] * [:EMAN2/Library:Python/C++ Programmers Documentation] * [:EMAN2/Concepts:Concepts in EMAN2] * [:EMAN2/Tutorials:EMAN2 Tutorials] * [:EMAN2/Galleries:Galleries] |
Please read this [[EMAN2/DatabaseWarning|Important Warning]] |
| Line 11: | Line 7: |
* [:EMAN2/FAQ:FAQ - Ask your questions here] |
* [[EMAN2/Install|Installation Instructions and Tips]] * User Documentation * [[EMAN2/Tutorials|EMAN2 Tutorials (START HERE!)]] * [[EMAN2/Programs|Individual Programs]] * [[EMAN2/Concepts|Concepts and Conventions in EMAN2]] * [[EMAN2/Parallel|Parallel Computing (multiple cores, linux clusters, sets of workstations)]] * [[EMAN2/Galleries|Galleries]] * Advanced Users & Programmers * [[EMAN2/Library|Python/C++ Programmers Documentation]] * [[EMAN2/FAQ|FAQ - Ask your questions here]] |
| Line 15: | Line 19: |
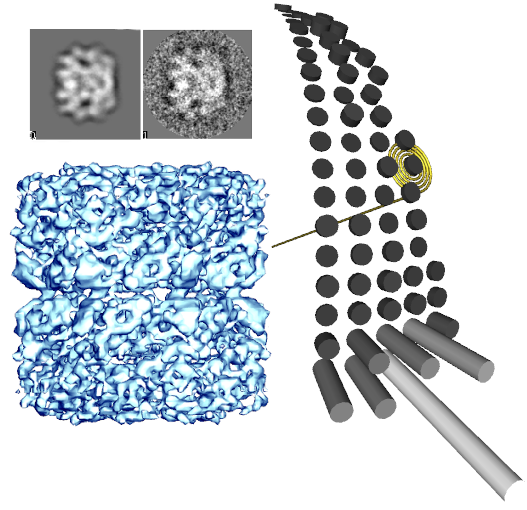
| EMAN2 is not (yet) a replacement for EMAN1, but can enhance EMAN1's capabilities. It was designed so both EMAN1 and EMAN2 can be installed in the same user account with no conflicts. All EMAN2 programs, including GUI programs are written in the easy-to-learn Python scripting language. This permits knowledgable end-users to customize any of the code with unprecendented ease. If you aren't an advanced user, you can still make use of all of EMAN2's command-line programs, all of which start with 'e2'. Any programs in EMAN2 with an EMAN1 equivalent are likely substantially improved over their EMAN1 equivalents. For example e2pdb2mrc.py is ~10x faster than the EMAN1 pdb2mrc. | EMAN2 is the successor to [[EMAN1|EMAN1]]. It is a broadly based greyscale scientific image processing suite with a primary focus on processing data from transmission electron microscopes. EMAN's original purpose was performing single particle reconstructions (3-D volumetric models from 2-D cryo-EM images) at the highest possible resolution, but the suite now also offers support for single particle cryo-ET, and tools useful in many other subdisciplines such as helical reconstruction, 2-D crystallography and whole-cell tomography. Image processing in a suite like EMAN differs from consumer image processing packages like Photoshop in that pixels in images are represented as floating-point numbers rather than small (8-16 bit) integers. In addition, image compression is avoided entirely, and there is a focus on quantitative analysis rather than qualitative image display. EMAN2 is now a fully usable reconstruction package, including initial parallelism support. However, it was designed so both EMAN1 and EMAN2 can be installed in the same user account with no conflicts, in case you need some EMAN1 functionality. All EMAN2 programs, including GUI programs are written in the easy-to-learn Python scripting language. This permits knowledgable end-users to customize any of the code with unprecendented ease. If you aren't an advanced user, you can still make use of all of EMAN2's command-line programs, all of which start with 'e2'. Any programs in EMAN2 with an EMAN1 equivalent are likely substantially improved over their EMAN1 equivalents. For example e2pdb2mrc.py is ~10x faster than the EMAN1 pdb2mrc. {{attachment:idea_5_c.png}} |
EMAN2
EMAN2 is now stable enough to be a complete replacement for EMAN1 for most tasks. It is capable of single particle reconstructions at near-atomic resolution, and can process very large data sets (>100,000 particles).
Please read this Important Warning
- User Documentation
Advanced Users & Programmers
About EMAN2
EMAN2 is the successor to EMAN1. It is a broadly based greyscale scientific image processing suite with a primary focus on processing data from transmission electron microscopes. EMAN's original purpose was performing single particle reconstructions (3-D volumetric models from 2-D cryo-EM images) at the highest possible resolution, but the suite now also offers support for single particle cryo-ET, and tools useful in many other subdisciplines such as helical reconstruction, 2-D crystallography and whole-cell tomography. Image processing in a suite like EMAN differs from consumer image processing packages like Photoshop in that pixels in images are represented as floating-point numbers rather than small (8-16 bit) integers. In addition, image compression is avoided entirely, and there is a focus on quantitative analysis rather than qualitative image display.
EMAN2 is now a fully usable reconstruction package, including initial parallelism support. However, it was designed so both EMAN1 and EMAN2 can be installed in the same user account with no conflicts, in case you need some EMAN1 functionality. All EMAN2 programs, including GUI programs are written in the easy-to-learn Python scripting language. This permits knowledgable end-users to customize any of the code with unprecendented ease. If you aren't an advanced user, you can still make use of all of EMAN2's command-line programs, all of which start with 'e2'. Any programs in EMAN2 with an EMAN1 equivalent are likely substantially improved over their EMAN1 equivalents. For example e2pdb2mrc.py is ~10x faster than the EMAN1 pdb2mrc.